Impact of Cathepsin L Inhibition in SARS-CoV-2 Infection and Potential Therapeutic Interventions
Muteeba Azhar1, Asma Irshad1*, Ansar Zubair2, Tahira Batool3, Ammara Maqsood4, Mehreen Saleem5, and Rida Bashir6
1School of Biochemistry and Biotechnology, University of the Punjab, Lahore, Pakistan
2Department of Zoology, University of Education, Lahore, Pakistan
3Allied Health Sciences, Superior University, Lahore, Pakistan
4Centre for Excellence in Solid State Physics, University of the Punjab, Lahore, Pakistan
5Department of Physiology, The Islamia University of Bahawalpur, Pakistan
6Shalamar Institute of Health Sciences, Shalamar Hospital, Lahore, Pakistan

Abstract
Background. The global COVID-19 pandemic caused by SARS-CoV-2 has prompted the urgent development of effective therapeutic strategies since its emergence in China. Cathepsin L is a lysosomal cysteine protease that plays a pivotal role in the entry of SARS-CoV-2 into the host cell. It follows an endocytic pathway that triggers the fusion of host and viral cell membranes.
Mechanism. Viral RNA is released during this phase and enters the host's cytoplasm through cleavage at S1/S2 or S2′ sites of the Spike glycoprotein of SARS-CoV-2. A study showed K790 as the potential cleavage site for cathepsin L. It is located near the S2′ site on the same loop. Its potential for proteolysis indicates its capacity to induce structural modifications analogous to S2′ cleavage, ultimately activating membrane fusion to allow the entry of the virus. The inhibitors of cathepsin L have emerged as effective drug targets in antiviral therapy.
Conclusion. This study aims to elaborate on the potential role of cathepsin L in SARS-CoV-2 infection during its entry into the host cells and also explores its functional and structural biology. Additionally, it highlights several promising inhibitory compounds including K11777, E64d, SDI 26681509, CID23631927, and Gallinamide A, which are effective in treating the SARS-CoV-2 infection. US Food and Drug Administration (FDA) approved drugs including amantadine, teicoplanin, and chloroquine have the potential to combat the SARS-CoV-2 infection. Understanding the significance of cathepsin L and the use of its inhibitors as therapeutic agents may open new opportunities for developing effective treatments for SARS-CoV-2.
GRAPHICAL ABSTRACT

Highlights
- Cathepsin L plays a pivotal role in SARS-CoV-2 infection by facilitating viral entry through the cleavage of the S glycoprotein.
- Structural and functional biology of cathepsin L provides insights into developing effective therapies against the SARS-CoV-2 infection.
- Promising inhibitors, such as K11777 and E64d and FDA-approved drugs, such as amantadine and chloroquine show efficacy in targeting cathepsin L to potentially block viral replication.
1. Introduction
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) was first identified in December 2019 as a viral disease in Wuhan, China, which rapidly spread globally [1]. On January 10, 2020, the first SARS-CoV-2 genome was made accessible to the public. The genome of SARS-CoV-2 is enveloped, possessing surface proteins that form spikes [2]. It comprises a positive-sense single-stranded RNA (ssRNA+) with a size ranging from 26 to 32 kilobases [1]. This virus shares more than 90% of its genetic makeup with SARS-CoV. The emergence of SARS-COV-2 variants necessitates real-time surveillance. Over 11 million sequences have been shared through GISAID by May 2022. Moreover, outbreak.info was used to track 40 million lineage-mutation combinations over 7,000 locations [3]. This virus is presumably more contagious than SARS-CoV and MERS-CoV. SARS-CoV-2 has given rise to several strains with varying degrees of severity and transmissibility, such as alpha (B.1.1.7), beta (B.1.351), gamma (P.1), delta (B.1.617.2), and Omicron (B.1.1.529). These strains were first identified in Brazil, the United Kingdom, India, and South Africa respectively and eventually became dominant worldwide in mid-2021 [4]. A total of 774,834,251 COVID-19 cases were confirmed by the World Health Organization (WHO) by March 3, 2024 including 7,037,007 deaths, worldwide. The number of vaccine doses administered to date totals 13,595,583,125. This information can be found in Figure 1 of the Epidemiology 2023 report on the WHO's COVID-19 website [5].

Figure 1. WHO COVID-19 Reported Cases 5
A previous study aimed to standardize the literature on the clinical manifestations of COVID-19 patients. In this regard, 26 different clinical manifestations were reported, out of which 6 symptoms had a prevalence of not less than 25%. These included fever, fatigue, dyspnea, cough, malaise, and sputum/secretion. Anorexia, myalgia, rhinitis, sneezing, sore throat, headache, chest pain, diarrhea, and neurological and dermatological disorders were among the other prevalent symptoms crucial for early diagnosis and preventive measures [6]. The respiratory system is the most frequently affected in individuals who experience clinical illness due to SARS-CoV-2. The virus is zoonotic and associated with lethal illnesses. Dyspnea, multifocal pneumonitis (which can rapidly lower blood oxygen saturation), systemic cytokine storm, multiple organ system failure, and death are all possible complications that patients may experience. It is very challenging to monitor and manage transmission because most people are asymptomatic carriers but have positive laboratory reports [6].
The virus affects extrapulmonary organs that contain its receptor proteins. Various studies showed that viral infection occurs when the virus attaches to angiotensin-converting enzyme-2 (ACE2) receptors found in kidneys, liver, colon, brain, heart, throat, vascular endothelial cells, and other tissues [7]. The pathogenesis of SARS-CoV-2 is regulated by many proteolytic enzymes, such as transmembrane serine protease-2 (TMPRSS2), furin, and cathepsin L which permits the virus to enter and replicate within cells. TMPRSS2 and proteases furin get stimulated and initiate spike glycoprotein cleavages [8, 9]. This change starts membrane fusion by making the S glycoprotein fusion competent. Multiple endocytic pathways are activated by cathepsin L, allowing SARS-CoV-2 to enter the cells [10].
Cathepsin L is a lysosomal cysteine protease that is only activated in an acidic medium [11]. Depending upon its location in the cell, it performs various physiological functions. It cleaves the S glycoprotein of the virus into its subunits (CS-1 and CS-2). All the discovered SARS-COV-2 variants have highly conserved cleavage sites [12]. Following this cleavage, host cells and viruses fuse their membranes together. Consequently, the viral genome (ssRNA) enters the host cell and begins its replication [13]. Cathepsin L inhibitors should be investigated as a therapeutic option due to the rapid development and spread of SARS-CoV-2 infection in individuals, globally [10]. It prevents SARS-CoV-2 entry into host cells and its intracellular proliferation without affecting the immune system [13]. To validate its effectiveness, more research is required. Various drugs and inhibitors show different levels of effectiveness in inhibiting cathepsin L. This review aims to discuss the mechanism of action of cathepsin L and explore how its inhibitors and drugs can be used to prevent viral entry into the host cells.
Functional Biology of Cathepsin L
Cathepsin L is a cysteine endopeptidase mainly found in lysosomes. It is encoded by the CTSL 1 gene in humans which is present on chromosome number 9 [14]. Its molecular weight is approximately 37,564 Da. In its spatial structure, it has an alpha helix in the L domain and beta sheets in the R domain [10]. Its activity and stability depend upon its ionic strength. It is catalytically active in an acidic medium with a pH range of 3.0-6.5 and an optimum pH of 5.5 [15]. To better understand its structure and interaction with inhibitors, the 3D structure of human cathepsin L bound to the covalent inhibitor E-64 (PDB ID: 8A4V 18) is rendered in this study using Chimera 1.17.3, as shown in Figure 2 [16].
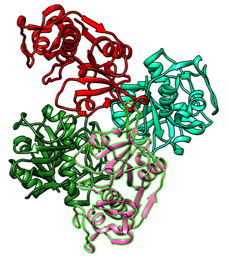
Figure 2. Human Cathepsin L with Covalently Bound E-64 [16]
Cathepsin L is also found in lysosomes, extracellular matrix, nucleus, and cytoplasm. It performs various physiological functions inside the cell [17]. When it is in lysosomes it is involved in antigen processing, immune response, growth, and development. Moreover, it also allows the entry of viruses into the cell. It performs apoptosis, necroptosis, and inflammation in the cytosol. Further, nuclear cathepsins control cell cycle progression and facilitate cell proliferation and differentiation. Cathepsin L in the extracellular matrix is involved in bone remodeling and EMC degradation [18]. Numerous human diseases including cancer, diabetes, obesity, cardiovascular and inflammatory illnesses, kidney problems, neurological disorders, as well as infectious diseases have been linked to a dysregulation of cathepsin expression or activity [17]. >
1.2. Mechanism of SARS-CoV-2 Entry
Several structural and nonstructural proteins in SARS-CoV-2 have a size of approximately 30 kb. The four major structural proteins namely spike (S), envelope (E), nucleocapsid (N), and membrane (M) expressed by structural genes are located before the 3′ end of the viral genome [19]. S glycoprotein holds the permit for the viral genome entry into the host cell by attaching to its membrane. The viral membrane has a crown-like appearance when assembled with several copies of the S protein as homotrimers. They enter the cell by two routes, that is, via membrane fusion or receptor-mediated endocytosis, depending upon the localization of peptidases [19].
The receptor binding domain (RBD) of S proteins is bound by human host cell receptors, allowing viral entry. The mechanism of SARS-CoV-2 entrance is primarily known to involve the angiotensin-converting enzyme 2 (ACE2) receptors. These receptors are abundantly present in testicular, kidney, liver, colon, lung, and vascular endothelial cells [20]. Some other host receptors promote virus invasion primarily into the respiratory system cells. After interacting with the RBD receptor, S glycoprotein is cleaved by proteases that include furin, TMPRSS2, and cathepsin B/L, which catalyze proteolytic cleavage [21]. Recently, two previously unidentified cleavage sites (CS-1 and CS-2) for the protease cathepsin L were identified in S protein [22]. Cathepsin L activates S proteins by cleaving them into its subunits S1 and S2. The binding subunit is S1, while the fusion subunit is S2 [23]. To release ssRNA into the host cytoplasm, the S2 subunit permits the viral membrane to fuse with the host cell. This viral genome now controls the machinery of the host cell [24].
1.3. Potential Role of Cathepsin L in SARS-CoV-2 Infection
The potential role of Cathepsin L is to allow the virus to penetrate the targeted host cells by activating S proteins that bind to it. When a virus infects a person, it allows membrane fusion, involving the binding of the receptor and the alteration of the S protein conformation [13]. In vitro SARS-CoV-1 membrane fusion has been demonstrated by prior research to rely on S protein proteolysis by host cathepsin L [25]. Consequently, it is feasible that medicines targeting cathepsin L can be developed to cure SARS-CoV-2 infection [10].
1.3.1. Cathepsin L Aids in Infection and Replication. According to research, the level of cathepsin L is high in people infected with SARS-CoV-2 and returns to normal after recovery from the infection [10]. It indicates that cathepsin L is essential for the penetration of SARS-CoV-2. Viral S protein undergoes conformational changes as a consequence of attachment to the receptor. Cathepsin L then cleaves the S protein in endosomes to cause the infection [26]. The surface of SARS-CoV-2 is characterized by a glycosylated trimer of mature S glycoprotein which is essential for its infection and replication. It has two subunits composed of approximately 1260 amino acids. The S1 subunit contains 685 amino acids that are divided into four domains: the receptor-binding domain (RBD) in the N-terminal domain (NTD), the C-terminal domain (CTD), and two subdomains (SD1 and SD2) [27]. The S2 subunit has 588 amino acids that form the stalk including a cytoplasmic tail (CT), transmembrane domain (TM), two heptad repeats (HR1 and HR2), and an N-terminal hydrophobic fusion peptide (FP) arranged as FP-HR1-HR2-TM-CT, as shown in Figure 3. Furin in the virus-producing cell cleaves SARS-CoV-2's S1-S2 border, but target-cell proteases are still needed to cleave the S2′ location. Cathepsin L and TMPRSS2 are involved in activating the S protein on plasma membrane and endolysosome, respectively [28].

Figure 3. Mechanism of SARS-CoV-2 entry
1.3.2. Cathepsin L Cleaves Spike Proteins. Cathepsin L cleaves the S protein to activate its fusion potential. The activated S protein regulates two cleavage processes. Initially, the S1 and S2 domains are cleaved apart. Afterward, a second cleavage at the S2 domain promotes the activation and unmasking of fusion peptides [26]. Cathepsin L cleaves the S protein between residues Thr696 and Met697 in the S1-S2 domains. This cleavage results in the release of the S1 subunit from the S2 subunit. The fusion peptide of the S2 subunit is inserted into the membrane of the endosome [29]. This causes conformational changes in S proteins. A six-helix bundle structure is formed when the S2 subunit folds back. This cleavage of the S protein increases the mediated cell-to-cell fusion by up to 70% [10]. All SARS-CoV-2 variations including the most recent variant Omicron required CTSL cleavage to be infected during a pseudo-virus (PsV) infection experiment [13].
Subsequently, viral and endosomal membranes interact and eventually fuse. The host cell cytoplasm is then invaded by the RNA released by coronaviridae. SARS-CoV-2 is a positive sense ssRNA virus with its own genetic material [14]. Without entering the nucleus, viral RNA can act as messenger RNA, instructing the ribosomes of the host cell to produce viral proteins. It has been proved that the process of spike activation is performed by cathepsin L through endocytosis, which is one potential method that causes infection [30].
To analyze the ability of cathepsin L to cleave the S protein of SARS-CoV-2, EV Matveev and his colleagues performed genome-wide bioinformatics analysis. They used UniProt and Protein Data Bank for amino acid sequencing and structural data retrieval and the SWISS-MODEL for loop modeling to investigate the viral S protein cleavage mechanisms.
SARS-CoV-2 mutations, genetic expressions, and protease localization are derived from various databases, such as COMPARTMENTS, TISSUES, and CoVariants. Positional-weighted matrices from the MEROPS database are used to construct protease sequence specificity models. Structural susceptibility to proteolysis is manifested by mapping proteolytic events to three-dimensional structures and by training a linear discriminant analysis model. The insights into SARS-CoV-2 cleavage sites and protease activation of S protein can also be obtained by using the aforementioned approach [31].
According to this research, the cleavage site (CS) value is observed as maximum at the K790 position which is close to S2′. Similar to S2′ cleavage, cleaving K790 may cause conformational changes that reveal fusion peptide and initiate membrane fusion. The structural suitability of K790 for proteolysis was evaluated by applying a bioinformatics technique to the three-dimensional structure of S proteins. K790 and R815 sites were the primary focus of the mapping of the S protein’s structural cleavage scores (SCSs) using the ACE2 receptor. The SCSs at both locations were in the loop next to the fusion peptide. They were suitably high and similar to 0.61 for K790 and 0.58 for R815. In comparison to other proteases at this position, cathepsin L exhibited the second-highest cleavage score. While mesotrypsin, exclusively expressed in the gastrointestinal tract, exhibited the highest score. In contrast, cathepsin L is present in various regions, including the respiratory system, gastrointestinal tract, and lungs. K790 is a novel cleavage site for cathepsin L, shedding light on the intricate mechanisms of SARS-CoV-2 entry and fusion activation. Moreover, robust validation supports K790 as a strong experimental candidate [31].
1.4. Inhibitors of Cathepsin L as Therapeutic Agents against SARS-COV-2
The inhibition of cathepsin L is a novel approach to combat SARS-CoV-2 infection. The scientific community became more interested in the development of cathepsin L inhibitors after the 2003 SARS outbreak [14]. The first cathepsin L inhibitor known as cystatin was isolated from Aspergillus in 1981 [32]. Recent research has revealed that cathepsin L and its newly identified cleavage sites within S proteins of SARS-CoV-2 are important targets for the development of therapies aimed at combating SARS-CoV-2 infection [12]. According to the research, the following compounds inhibited cathepsin L and showed anti-coronavirus activity [14, 32].
1.4.1. K11777. K11777 is a small molecule that is a potent and irreversible inhibitor of cathepsin L. It is a dipeptide analog that features an electrophilic vinyl-sulfone group and remains highly effective at covalently inhibiting cathepsins [32, 33].The molecule binds to the catalytic site of cathepsin L via the vinyl sulfone moiety. It serves as a Michael acceptor and combines with the Cys-SH residue in the active site to permanently inhibit the protease activity [34]. Targeted and potent inhibitory effects on the entry of SARS-CoV and Ebola virus show broad-spectrum antiviral activity by focusing on cathepsin-mediated viral entry into host cells [35]. K11777 has the potential to be used as viral infection treatment, as evidenced by phase 1 clinical trials that demonstrated that it can significantly minimize the infectivity of SARS-CoV-2 in a variety of host cells. K11777 inhibits the processing of S protein by cathepsin L and has been found in various mammalian cell lines to play a potent role in SARS-CoV-2 infection [36].
1.4.2. E64d. E-64d is a natural inhibitor isolated from Aspergillus japonicus in 1978. This cysteine protease inhibitor targets the enzyme cathepsin L and has been used in various studies [37]. This irreversible epoxysuccinyl inhibitor can enter cells and block viral entry [35]. It can alter the viral fusion process that occurs at the endosome membrane, which is essential for the entry of viruses into host cells [12]. Recent research suggests E64d as a potential treatment for SARS-CoV-2 infection in human cells and mice models, as it effectively inhibits the Omicron variant and reduces cathepsin L activity. Moreover, E-64d has been shown to block the entry of virus uptake by 92.5%, demonstrating its potential as a treatment for viral infections [38].
1.4.3. SDI 26681509. SID 26681509 is a small-molecule inhibitor belonging to the thiocarbazate class of compounds [38]. It has been found in specific targeting of cathepsin L, causing the blockage of SARS-CoV-2 entry into human embryonic kidney 293/hACE2 cells. This inhibitor SID 26,681,509 is strong and reversible and can reduce viral entry by 76% [35]. These findings suggest that SID 26681509 holds promise as a therapeutic drug in treating viral infections.
1.4.4 CID23631927. CID23631927 is a small-molecule tetrahydroquinoline oxocarbazate inhibitor that selectively and potently inhibits SARS-CoV by targeting the cathepsin L enzyme in a cellular context [10, 34]. It has been investigated as a human cathepsin L inhibitor and has potential as an antiviral agent against SARS-CoV and Ebola pseudo-type virus. Research showed that the cathepsin L specific inhibitor CID 23631927 can inhibit SARS-CoV entry into human embryonic kidney 293T cells, indicating its potential for therapeutic use [35].
1.4.5. Gallinamide A. Gallinamide A is referred to as somatostatin 4. It is a natural marine product discovered independently of the Schizothrix genus of marine cyanobacteria in Panama [39]. Gallinamide A was first identified by Miller and his colleagues as a powerful irreversible cathepsin L inhibitor with an IC50 value of 5 nM [40]. Gallinamide A and its analogs thus offer a wonderful inhibitory framework that may be used to create selective enzyme inhibitors for a range of therapeutic applications [41]. In vitro studies showed that Gallinamide A, along with two highly active natural product analogs that inhibit cathepsin L, are potent inhibitors of SARS-CoV-2 infection. Their respective EC50 values were found to be in the nanomolar range [42]. A recent study also showed that Gallinamide A has two potential analogs 15 and 16. They can prevent the onset of SARS-CoV-2 infection and overexpress cathepsin L at nanomolar concentrations in Vero E6 cells. Their respective EC50 values were found to be 28, 168, and 920 nM, while their CC50 values were greater than 100 μM [43].
1.5. FDA-APPROVED DRUGS
1.5.1. Amantadine. The US FDA approved amantadine in 1968 for the treatment of influenza and later for the treatment of Parkinson's disease [10]. Being a lysosomotropic drug, it disrupts the lysosomal pathway and lessens the likelihood of the spread of the virus. Drug screens with a high throughput are carried out to look for potential treatments. Smieszek and colleagues used this method to show that amantadine hydrochloride inhibits the expression of the mRNA of cathepsin L, possibly altering its functional environment [43]. Recent studies have demonstrated that amantadine effectively inhibits cathepsin L activity and prevents SARS-CoV-2 infection in humanized mice, thereby indicating that cathepsin L inhibitors have the potential to be used as therapy for SARS-CoV-2 infection [10, 26]. The use of amantadine may cause an increase in endosome pH and a decrease in cathepsin L expression. Additionally, in vitro studies conducted on the Vero 6 cell line have demonstrated that SARS-CoV-2 replication is inhibited by amantadine due to the blocking of viroporin [44].
1.5.2. Teicoplanin. Teicoplanin, along with its derivatives such as dalbavancin, oritavancin, and telavacin, have been identified as potential inhibitors of cathepsin L after undergoing high-throughput screening of FDA-approved drugs. Teicoplanin is a glycopeptide antibiotic in which the conservation of cathepsin L cleavage sites has been observed. Recent studies showed that teicoplanin can prevent the entrance of the pseudoviruses that cause MERS-CoV, SARS-CoV, SARS-CoV-2, and Ebola [45]. A Chinese study showed teicoplanin as able to dose-dependently prevent SARS-CoV-2 infection in A549, HEK293 T, and huh7 cells [13].
1.5.3. Chloroquine. Hydroxychloroquine/chloroquine has antiviral, immunomodulatory, and antithrombotic effects and may inhibit cathepsin L in preclinical studies. Commonly prescribed alone in COVID-19 treatment trials, it improves patients’ conditions in intensive care [46]. The combination of hydroxychloroquine, favipiravir, darunavir, and ritonavir reduced mortality by 25% as compared to the combination of oseltamivir, lopinavir, and ritonavir, with efficacy attributed to viral cycle inhibition [47]. A systematic review found that the FDA revoked emergency use authorization for hydroxychloroquine and chloroquine for COVID-19 treatment, requiring large clinical trials to determine efficacy and safety [48].
1.5.4. OtherTherapeutic Targets. Remdesivir is an antiviral drug that was first approved by FDA for emergency use as a cure for hospitalized patients tackling severe SARS-CoV-2 infection [49]. Cathepsin L activity can be inhibited by antimicrobial medication, such as teicoplanin, which is an antibiotic. Rifampicin is an anti-tuberculous medicine, while clofazimine has been identified as an anti-leprosy drug, and saquinavir is an anti-HIV drug that inhibits its activity [50]. Rifampicin has been reported to inhibit cathepsin L competitively and clofazimine as a non-competitive inhibitor [14]. Cathepsin L is inhibited by immunomodulatory medications, such as dexamethasone, which reduces its activity in muscle cells [14]. A recent study revealed that there are five molecules, namely Mg-132, Mg-101, leupeptin hemisulfate, Z-FA-FMK, and calpeptin which demonstrated potent inhibition of cathepsin L activity in the nanomolar range and impeded both live and pseudotype SARS-CoV-2 infection. Remarkably, scientists revealed daptomycin (an FDA-approved antibiotic) that exhibits a significant cathepsin L inhibitory effect which can reduce the spread of SARS-CoV-2 pseudovirus [51]. In a recent study, Sven Falke and colleagues characterized carbonyl- and succinyl epoxide-based inhibitors as anti-SARS-CoV-2 inhibitors. Calpain inhibitor namely XII and MG-101 and cathepsin L inhibitor IV show low EC50 activity in Vero E6 cells. Crystal structures of cathepsin L in complex with 14 compounds provide a basis for drug development [52].
1.5.5. Combine Therapeutic Effects. Combining cathepsin L inhibitors with other antiviral medicines is a viable technique to increase therapeutic effectiveness against SARS-CoV-2. This method makes use of complementary modes of action, which may result in synergistic effects that boost the total antiviral efficacy [53]. For example, combining cathepsin L inhibitors such as K11777 or E64d with known antiviral medicines such as remdesivir or favipiravir might target various phases of the viral life cycle, effectively lowering viral load more than monotherapy alone. Furthermore, this combination may increase the range of efficacy against developing SARS-CoV-2 variants that are resistant to single-agent therapy. Lower doses of each medicine in combination treatment may reduce adverse effects, enhancing patient tolerance and adherence [54].
2. CHALLENGES AND LIMITATIONS
Targeting cathepsin L as a treatment method against SARS-CoV-2 involves a number of hurdles, including specificity, possible side effects, and balancing efficacy and safety. Cathepsin L is involved in a number of physiological processes, including immunological response and antigen processing; therefore, specificity is an important consideration. Inhibiting this protease may impair these critical processes, resulting in unexpected effects in the host. Cathepsin L is widely expressed in several tissues; hence, systemic inhibition may have a negative impact on organs where Cathepsin L is important, such as the lungs and gastrointestinal tract. This needs a detailed assessment of the safety profile of all Cathepsin L inhibitors. Furthermore, establishing a balance between efficacy and toxicity is crucial. While cathepsin L inhibitors may successfully block viral entry, their therapeutic window must be determined precisely. Combining these inhibitors with other antiviral medications may improve overall efficacy while decreasing adverse effects, although thorough clinical testing is necessary to optimize therapy regimens. Addressing these issues is critical to establish effective and safe treatment strategies targeting cathepsin L in the battle against COVID-19.
Future study into cathepsin L's role in SARS-CoV-2 infection would fill many crucial gaps. To begin with, a better knowledge of how cathepsin L interacts with viral proteins other than S glycoprotein is required to determine its complete influence on viral entrance and reproduction. Moreover, studying the impact of various SARS-CoV-2 variants on cathepsin L activity may reveal possible resistance pathways and influence treatment methods. Furthermore, long-term research is required to determine the physiological effects of cathepsin L inhibition on host immunological responses and general health. Finally, investigating the synergistic effects of combining cathepsin L inhibitors with other antiviral drugs may increase therapeutic efficacy while decreasing the possibility of resistance, opening the pathway for more effective COVID-19 therapies.
3. CONCLUSION
The study demonstrates numerous possible inhibitors, including K11777 and FDA-approved medications such as amantadine and chloroquine, that have the ability to disrupt the mechanism facilitating viral entry. Understanding the mechanisms of cathepsin L and its inhibitors may lead to novel antiviral therapeutics for COVID-19, addressing important hurdles in treatment effectiveness and safety. Future research should focus on optimizing these inhibitors and developing combination medicines to improve their efficacy.
Conflict of Interest
The author of the manuscript has no financial or non-financial conflict of interest in the subject matter or materials discussed in this manuscript.
Data Availability Statement
The data associated with this study will be provided by the corresponding author upon request.
Funding details
This research did not receive grant from any funding source or agency.
Bibliography
- Lai C-C, Shih T-P, Ko W-C, Tang H-J, Hsueh P-R. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and coronavirus disease-2019 (COVID-19): the epidemic and the challenges. Int J Antimicrob Agents. 2020;55(3):e105924. https://doi.org /10.1016/j.ijantimicag.2020.105924
- Pal M, Berhanu G, Desalegn C, Kandi V. Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2): an update. Cureus. 2020;12(3):e7423. https://doi.org/10.7759/cureus.7423
- Gangavarapu K, Latif AA, Mullen JL, et al. Outbreak. info genomic reports: scalable and dynamic surveillance of SARS-CoV-2 variants and mutations. Nat Methods. 2023;20(4):512–522. https://doi.org/10.1038/s41592-023-01769-3
- Zahmatkesh S, Sillanpaa M, Rezakhani Y, Wang C. Review of concerned SARS-CoV-2 variants like Alpha (B. 1.1. 7), Beta (B. 1.351), Gamma (P. 1), Delta (B. 1.617. 2), and Omicron (B. 1.1. 529), as well as novel methods for reducing and inactivating SARS-CoV-2 mutants in wastewater treatment facilities. J Hazard Mater Adv. 2022:e100140. https://doi.org/10.1016/j.hazadv.2022.100140
- Word Health Organization. Number of COVID-19 cases reported to WHO. World Health Organization Web site. https://data.who.int/dashboards/covid19/cases?n=c. Accessed March 3, 2024.
- da Rosa Mesquita R, Francelino Silva Junior LC, Santos Santana FM, et al. Clinical manifestations of COVID-19 in the general population: systematic review. Wien Klin Wochenschr. 2021;133(7-8):377–382. https://doi.org/10.1007/s00508-020-01760-4
- Gupta A, Madhavan MV, Sehgal K, et al. Extrapulmonary manifestations of COVID-19. Nat Med. 2020;26(7):1017–1032. https://doi.org/10.1038/s41591-020-0968-3
- Essalmani R, Jain J, Susan-Resiga D, et al. Distinctive roles of furin and TMPRSS2 in SARS-CoV-2 infectivity. J Virol. 2022;96(8):e00128-22. https://doi.org/10.1128/jvi.00128-22
- Zhao M-M, Yang W-L, Yang F-Y, et al. Cathepsin L plays a key role in SARS-CoV-2 infection in humans and humanized mice and is a promising target for new drug development. Sig Transduct Target Ther. 2021;6(1):e134. https://doi.org/10.1038/s41392-021-00558-8
- Neerukonda S. Therapeutic targets of host factors for potential COVID-19 treatments. In: Jen-Tsung Chen J-T, ed. Bioactive Compounds Against SARS-CoV-2. CRC Press; 2024:23–38.
- Kondo Y, Rajapakse S, Ogiwara K. Involvement of cathepsin L in the degradation and degeneration of postovulatory follicle of the medaka ovary. Biol Reproduct. 2023;109(6):904–917. https://doi.org/10.1093/biolre/ioad116
- Zhao M-M, Zhu Y, Zhang L, et al. Novel cleavage sites identified in SARS-CoV-2 spike protein reveal mechanisms for cathepsin L-facilitated viral infection and treatment strategies. Cell Discov. 2022;8(1):e53. https://doi.org/10.1038/s41421-022-00419-w
- Liu T, Luo S, Libby P, Shi G-P. Cathepsin L-selective inhibitors: a potentially promising treatment for COVID-19 patients. Pharmacol Therapeut. 2020;213:e107587. https://doi.org/10.1016/j.pharmthera.2020.107587
- Gaikwad P. Regulation of Cathepsin L Expression And Activity By Cell Confluence And The Circadian Clock [doctoral dissertation]. Fairborn, United States: Wright State University; 2023.
- Reinheckel T, Tholen M. Low‐level lysosomal membrane permeabilization for limited release and sublethal functions of cathepsin proteases in the cytosol and nucleus. FEBS Open Bio. 2022;12(4):694–707. https://doi.org /10.1002/2211-5463.13385
- Falke S, Lieske J, Guenther S, et al. Crystal Structure Of Human Cathepsin L With Covalently Bound E-64. Helmholtz Association; 2024. https://doi.org/10.2210/pdb8A4V/pdb
- Berdowska I, Matusiewicz M. Cathepsin L, transmembrane peptidase/serine subfamily member 2/4, and other host proteases in COVID-19 pathogenesis–with impact on the gastrointestinal tract. World J Gastroenterol. 2021;27(39): 6590–6600. https://doi.org/10.3748 /wjg.v27.i39.6590
- Yadati T, Houben T, Bitorina A, Shiri-Sverdlov R. The ins and outs of cathepsins: physiological function and role in disease management. Cells. 2020;9(7):e1679. https://doi.org/ 10.3390/cells9071679
- Yadav R, Chaudhary JK, Jain N, et al. Role of structural and non-structural proteins and therapeutic targets of SARS-CoV-2 for COVID-19. Cells. 2021;10(4):e821. https://doi.org/ 10.3390/cells10040821
- Gheblawi M, Wang K, Viveiros A, et al. Angiotensin-converting enzyme 2: SARS-CoV-2 receptor and regulator of the renin-angiotensin system: celebrating the 20th anniversary of the discovery of ACE2. Circulat Res. 2020;126(10):1456–1474. https://doi.org/10.1161/CIRCRESAHA.120.317015
- Bestle D, Heindl MR, Limburg H, et al. TMPRSS2 and furin are both essential for proteolytic activation of SARS-CoV-2 in human airway cells. Life Sci Allian. 2020;3(9):e202000786. https:// doi.org/10.26508/lsa.202000786
- Yang J-K, Zhao M-M, Zhu Y, et al. Novel Sites For Cathepsin L Cleavage in SARS-CoV-2 Spike Guide Treatment Strategies. Research Sequare; 2021. https://doi.org/10 .21203/rs.3.rs-734963/v1
- Jaimes JA, Millet JK, Whittaker GR. Proteolytic cleavage of the SARS-CoV-2 spike protein and the role of the novel S1/S2 site. iScicence. 2020;23(6):e101212. https://doi.org/ 10.1016/j.isci.2020.101212
- Zhang Q, Xiang R, Huo S, et al. Molecular mechanism of interaction between SARS-CoV-2 and host cells and interventional therapy. Signal Transduct Target Ther. 2021;6(1):e233. https://doi.org/10. 1038/s41392-021-00653-w
- Simmons G, Gosalia DN, Rennekamp AJ, Reeves JD, Diamond SL, Bates P. Inhibitors of cathepsin L prevent severe acute respiratory syndrome coronavirus entry. Proc Nat Acad Sci. 2005;102(33):e11876–11881. https://doi.org/10.1073/pnas.0505577102
- Gomes CP, Fernandes DE, Casimiro F, et al. Cathepsin L in COVID-19: from pharmacological pieces of evidence to genetics. Front Cell Infect Microbiol. 2020;10:e589505. https://doi.org/ 10.3389/fcimb.2020.589505
- Schroeder JT, Bieneman AP. The S1 subunit of the SARS-CoV-2 spike protein activates human monocytes to produce cytokines linked to COVID-19: relevance to Galectin-3. Front Immunol. 2022;13;831763. https://doi.org/10.3389/fimmu.2022.831763
- Jackson CB, Farzan M, Chen B, Choe H. Mechanisms of SARS-CoV-2 entry into cells. Nat Rev Molecul Cell Biol. 2022;23(1):3–20.
- Yu S, Zheng X, Zhou B, et al. SARS-CoV-2 spike engagement of ACE2 primes S2′ site cleavage and fusion initiation. Proc Nat Acad Sci. 2022;119(1):e2111199119. https://doi.org/10.1073/pnas.2111199119
- Ou X, Liu Y, Lei X, et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun. 2020;11(1):e1620. https://doi.org/10.1038/s41467-020-15562-9
- Matveev EV, Ponomarev GV, Kazanov MD. Genome-wide bioinformatics analysis of human protease capacity for proteolytic cleavage of the SARS-CoV-2 spike glycoprotein. Microbiol Spectr. 2024;12(2):e03530-23. https://doi.org/ 10.1128/spectrum.03530-23
- Li Q, Zhou S-R, KIM H, Wang H, Zhu J-J, Yang J-K. Discovering novel cathepsin L inhibitors from natural products using artificial intelligence. Comput Struct Biotechnol J. 2024;23:2606–2614. https://doi.org/ 10.1016/j.csbj.2024.06.009
- Mellott DM, Tseng C-T, Drelich A, et al. A Cysteine Protease Inhibitor Blocks SARS-CoV-2 Infection of Human and Monkey Cells. bioRxive. https://doi.org/10.1101/2020.10.23.347534
- Cannalire R, Stefanelli I, Cerchia C, Beccari AR, Pelliccia S, Summa V. SARS-CoV-2 entry inhibitors: small molecules and peptides targeting virus or host cells. Int J Mol Sci. 2020;21(16):e5707. https://doi.org/ 10.3390/ijms21165707
- Pislar A, Mitrović A, Sabotič J, et al. The role of cysteine peptidases in coronavirus cell entry and replication: the therapeutic potential of cathepsin inhibitors. PLoS Pathog. 2020;16(11):e1009013. https://doi.org /10.1371/journal.ppat.1009013
- Mellott DM, Tseng C-T, Drelich A, et al. A clinical-stage cysteine protease inhibitor blocks SARS-CoV-2 infection of human and monkey cells. ACS Chem Biol. 2021;16(4):642–650. https://doi.org/10.1021/acschembio.0c00875
- Chang M-C, Chen J-H, Lee H-N, et al. Inducing cathepsin L expression/production, lysosomal activation, and autophagy of human dental pulp cells by dentin bonding agents, camphor quinone, and BisGMA and the related mechanisms. Biomater Adv. 2023;145:e213253. https://doi.org/10.1016/j.bioadv.2022.213253
- Acacio Santini Pereira B, Souza-Silva F, Silva-Almeida M, et al. Proteinase inhibitors: a promising drug class for treating leishmaniasis. Curr Drug Targets. 2014;15(12):1121–1131.
- Ashhurst AS, Tang AH, Fajtova P, et al. Potent in vitro anti-SARS-CoV-2 activity by gallinamide A and analogs via inhibition of cathepsin L. bioRxiv. 2020:e2020.12.23.424111. https://doi. org/10.1101/2020.12.23.424111
- Miller B, Friedman AJ, Choi H, et al. The marine cyanobacterial metabolite gallinamide A is a potent and selective inhibitor of human cathepsin L. J Nat Prod. 2014;77(1):92–99. https://doi.org/10.1021/np400727r
- Dana D, Pathak SK. A review of small molecule inhibitors and functional probes of human cathepsin L. Molecules. 2020;25(3):e698. https:// doi.org/10.3390/molecules25030698
- Ashhurst AS, Tang AH, Fajtová P, et al. Potent anti-SARS-CoV-2 activity by the natural product gallinamide A and analogs via inhibition of cathepsin L. J Med Chem. 2021;65(4):2956–2970. https://doi.org/10.1021/acs .jmedchem.1c01494
- Smieszek SP, Przychodzen BP, Polymeropoulos MH. Amantadine disrupts lysosomal gene expression: a hypothesis for COVID-19 treatment. Int J Antimicrob Agents. 2020;55(6):e106004. https://doi.org/ 10.1016/j.ijantimicag.2020.106004
- Toft-Bertelsen TL, Jeppesen MG, Tzortzini E, et al. Amantadine inhibits known and novel ion channels encoded by SARS-CoV-2 in vitro. Commun Biol. 2021;4(1):e1347. https://doi.org/ 10.1038/s42003-021-02866-9
- Schütz D, Ruiz-Blanco YB, Münch J, Kirchhoff F, Sanchez-Garcia E, Müller JA. Peptide and peptide-based inhibitors of SARS-CoV-2 entry. Adv Drug Deliver Rev. 2020;167:47–65. https://doi.org/10.1016/j.addr.2020.11.007
- Roldan EQ, Biasiotto G, Magro P, Zanella I. The possible mechanisms of action of 4-aminoquinolines (chloroquine/hydroxychloroquine) against Sars-Cov-2 infection (COVID-19): a role for iron homeostasis? Pharmacol Res. 2020;158:e104904. https://doi.org/10.1016/j.phrs.2020.104904
- Jaimes-Castelán EG, González-Espinosa C, Magos-Guerrero GA, et al. Drugs and natural products for the treatment of COVID-19 during 2020, the first year of the pandemic. Bol Med Hosp Infant Mex. 2024;81(1):53–72. https://doi.org/10.24875/bmhim.23000016
- Babaei F, Mirzababaei M, Nassiri‐Asl M, Hosseinzadeh H. Review of registered clinical trials for the treatment of COVID‐19. Drug Develop Res. 2021;82(4):474–493. https://doi.org/10.1002/ddr.21762
- Awadasseid A, Wu Y, Tanaka Y, Zhang W. Effective drugs used to combat SARS-CoV-2 infection and the current status of vaccines. Biomed Pharmacother. 2021;137:e111330. https://doi.org/10.1016/j.biopha.2021.111330
- Yousefi B, Valizadeh S, Ghaffari H, Vahedi A, Karbalaei M, Eslami MJ. Global treatments for coronaviruses including COVID‐19. J Cell Phy. 2020;235(12):9133–9142. https://doi.org/10.1002/jcp.29785
- Yang W-L, Li Q, Sun J, et al. Potential drug discovery for COVID-19 treatment targeting Cathepsin L using a deep learning-based strategy. Comput Struct Biotechnol J. 2022;20:2442–2454. https://doi.org/ 10.1016/j.csbj.2022.05.023
- Falke S, Lieske J, Herrmann A, et al. Structural elucidation and antiviral activity of covalent Cathepsin L inhibitors. J Med Chem. 2024;67(9):7048–7067. https://doi.org /10.1021/acs.jmedchem.3c02351
- Milan Bonotto R, Mitrović A, Sosič I, et al. Cathepsin inhibitors nitroxoline and its derivatives inhibit SARS-CoV-2 infection. Antiviral Res. 2023;216:e105655. https://doi.org/10 .1016/j.antiviral.2023.105655
- Bobrowski T, Chen L, Eastman RT, et al. Synergistic and antagonistic drug combinations against SARS-CoV-2. Molecul Ther: J Am Soc Gene Ther. 2021;29(2):873–885. https://doi.org/10.1016/j.ymthe.2020.12.016
