Whole Genomic Sequencing and Analysis of Bacillus subtilis PWA: Linking Plastic Bioconversion to PHA Production
Abstract
 Abstract Views: 34
Abstract Views: 34
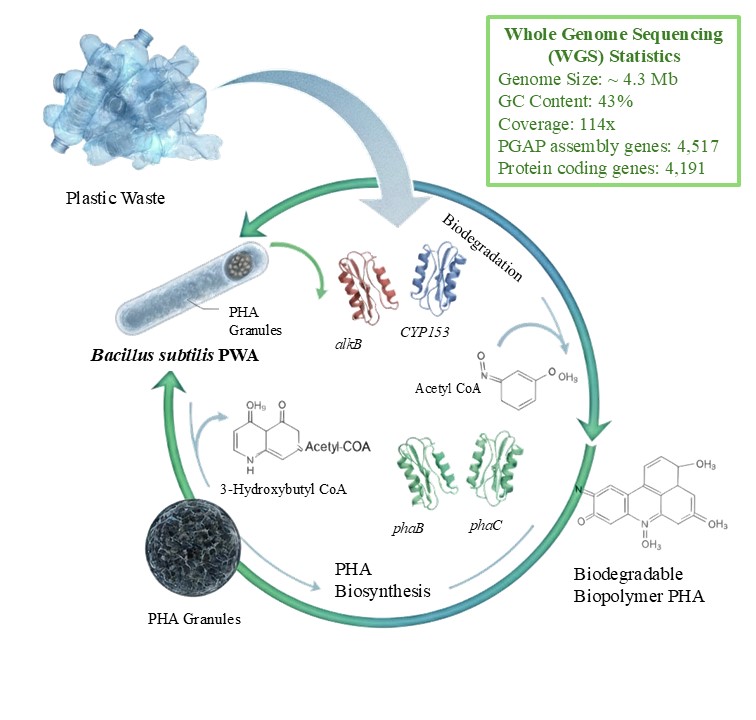
Plastic pollution requires innovative microbial solutions for plastic valorisation into sustainable bioproducts, such as intracellular PHA, during stressed conditions. This study reports the whole genome sequencing (WGS) and annotation of Bacillus subtilis PWA, using plastics for biopolymer PHA production. High throughput genome sequencing using Illumina NovaSeq 6000 gave a total of 983,667 raw reads with 114x average genome coverage. Genome assembly (4.3 Mb draft genome with 43.18% GC content) was submitted to NCBI database under the assembly number GCA_047314825.1. Approximately 4,517 genes (4,191 protein-coding) were annotated by NCBI Prokaryotic Genome Annotation Pipeline (PGAP). Functional annotation revealed genes associated with plastic degradation (alkB1, alkB2, alkB3, and CYP153) and PHA biosynthesis (phaA, phaB, phaC, phaJ, phaZ). The identification of genes concerned with survival and adaptation, for instance, osmoprotectant uptake (opuAA, opuAB, opuCA, opuCC), betaine biosynthesis (betB, opuD), and choline metabolism (natAB) supports the ability of strain PWA to survive in extremely stressed environments. Genome was visualized using Proksee CGViewer, while protein structure was visualized using AlphaFold. Metabolic pathways studied in the Kyoto Encyclopaedia of Genes and Genomes (KeGG) demonstrate modules for hydrocarbon degradation and PHA biosynthesis. The findings of this study including the genomic repertoire underscore the circular bioeconomic potential of the strain PWA for large-scale bioplastic production through the bioconversion of plastic wastes. Future studies should focus on metabolic engineering of PHA operon for optimized and volumetric production.
Downloads
References
Abid S, Raza ZA, Hussain T. Production kinetics of polyhydroxyalkanoates by using Pseudomonas aeruginosa gamma ray mutant strain EBN-8 cultured on soybean oil. 3 Biotech. 2016;6(2):e142. https://doi.org/10.1007/s13205-016-0452-4
Adeleke BS, Ayangbenro AS, Babalola OO. Genomic analysis of endophytic Bacillus cereus T4S and its plant growth-promoting traits. Plants. 2021;10(9):e1776. https://doi.org/10.3390/plants10091776
Alvarez-Santullano N, Villegas P, Mardones MS, et al. Genome-wide metabolic reconstruction of the synthesis of polyhydroxyalkanoates from sugars and fatty acids by Burkholderia sensu lato species. Microorganisms. 2021;9(6):e1290. https://doi.org/10.3390/ microorganisms9061290
Modi A, Vai S, Caramelli D, Lari M. The Illumina sequencing protocol and the NovaSeq 6000 system. In: Mengoni A, Bacci G, Fondi M, eds. Bacterial Pangenomics: Methods and Protocols. Springer; 2021:15-42.
Li W, Shen S, Wang J. Whole Genome sequencing assembly and annotation of halobacillus trueperi S61 isolated from the Qarhan Salt Lake. Resesracah Sequare; 2022. https://doi.org/10.21203/rs.3.rs-1188620/v1
Cao H, Peng T, Zhao W, Huang H, Yu S, Zhu Y. Whole-genome sequencing uncovers the plant growth-promoting potential of Bacillus licheniformis G41, isolated from the rhizosphere soil of Gannan navel orange. Ann Microbiol. 2025;75(1):e8. https://doi.org/10.1186/s13213-025-01797-8
Bankevich A, Nurk S, Antipov D, et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. 2012;19(5):455-477. https://doi.org/10.1089/ cmb.2012.0021
Blin K, Shaw S, Vader L, et al. antiSMASH 8.0: extended gene cluster detection capabilities and analyses of chemistry, enzymology, and regulation. Nucleic Acids Res, 2025;53(7):W32-W38. https:// doi.org/10.1093/nar/gkaf334
Dergham Y, Sanchez-Vizuete P, Le Coq D, et al. Comparison of the genetic features involved in Bacillus subtilis biofilm formation using multi-culturing approaches. Microorganisms. 2021;9(3):e633. https://doi.org/10.3390/microorganisms9030633
Langille MG, Zaneveld J, Caporaso JG, et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol. 2013;31(9):e814. https://doi.org/ 10.1038/nbt.2676
Mishra R, Chavda P, Kumar R, et al. Exploring genetic landscape of low-density polyethylene degradation for sustainable troubleshooting of plastic pollution at landfills. Sc Total Environ. 2024;912:e168882. https://doi.org/10.1016/j.scitotenv.2023.168882
Narancic T, Salvador M, Hughes GM, et al. Genome analysis of the metabolically versatile Pseudomonas umsongensis GO16: the genetic basis for PET monomer upcycling into polyhydroxyalkanoates. Microbial Biotechnol. 2021;14(6):2463-2480. https://doi.org/10. 1111/1751-7915.13712
Fedorova ND, Moktali V, Medema MH. Bioinformatics approaches and software for detection of secondary metabolic gene clusters. In: Keller NP, Turner G, eds. Fungal Secondary Metabolism: Methods and Protocols. 2012:23-45. https://doi.org/10.1007/978-1-62703-122-6_2
Sohail R, Batool R, Jamil N. PHA synthesis by bacteria using low density polyethylene, starches and cellulosics. Korean J Microbiol. 2021;57(3):183-196. https://doi.org/10.7845/kjm.2021.1012
Canham J, Win J, Network G, Kamoun S. GetGenome: overcoming inequalities in access to genomics technology. PLoS Biol. 2024;22(8):e3002804. https://doi.org/10.1371/journal.pbio.3002804
Chau C, Mohanan G, Macaulay I, Actis P, Wälti C. Automated purification of DNA origami with SPRI beads. Small. 2024;20(20):e2308776. https://doi.org/10.1002/smll.202308776
Stortchevoi A, Levine SS. Improved Yield of SPRI Beads–Based size selection in the very high molecular weight range. J Biomolecul Tech. 2024;35(1):e3fc1f5fe.d13e7666. https://doi.org/10.7171/3fc1f5fe.d13e7666
Dang J, Mendez P, Lee S, et al. Development of a robust DNA quality and quantity assessment qPCR assay for targeted next-generation sequencing library preparation. Int J Oncol. 2016;49(4):1755-1765. https://doi.org/10.3892/ijo.2016.3654
Handžić N, Pećar D, Durgut S, et al. Optimization of Illumina® Nextera™ XT library preparation for whole mitochondrial genome sequencing and confirmatory Sanger sequencing. Medicinski Glasnik. 2025;22(2):e191. https://doi.org/10.17392/1950-22-02
Fan L. HSPA-A universal graphical user interface for the Hamilton Microlab STAR liquid handler. J Appl Bioanal. 2016;2(1):38-44. http://dx.doi.org/10.17145/jab.16.006
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30(15):2114-2120. https://doi.org/10.1093/bioinformatics/btu170
Seemann T. Prokka: rapid prokaryotic genome annotation. Bioinformatics. 2014;30(14):2068-2069. https://doi.org/10.1093/bioinformatics/btu153
Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 2018;35(6):1547-1549. https://doi.org/10.1093/molbev/msy096
Tatusova T, DiCuccio M, Badretdin A, et al. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016;44(14):6614-6624. https://doi.org/10.1093/nar/gkw569
Kanehisa M, Furumichi M, Sato Y, Matsuura Y, Ishiguro-Watanabe M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 2025;53(D1):D672-D677. https://doi.org/10.1093/nar/gkae909
Grant JR, Enns E, Marinier E, et al. Proksee: in-depth characterization and visualization of bacterial genomes. Nucleic Acids Res. 2023;51(W1):W484-W492. https://doi.org/10.1093/nar/gkad326
Varadi M, Bertoni D, Magana P, et al. AlphaFold protein structure database in 2024: providing structure coverage for over 214 million protein sequences. Nucleic Acids Res. 2024;52(D1):D368-D375. https://doi.org/10.1093/nar/gkad1011
Wafaa DM, Sadik MW, Eissa HF, Tonbol K. Biodegradation of low-density polyethylene LDPE by marine bacterial strains Gordonia alkanivorans PBM1 and PSW1 isolated from Mediterranean Sea, Alexandria, Egypt. Sci Rep. 2025;15(1):1-15. https://doi.org/10.1038/s41598-025-96811-z
Alotaibi F, St-Arnaud M, Hijri M. In-depth characterization of plant growth promotion potentials of selected alkanes-degrading plant growth-promoting bacterial isolates. Front Microbiol. 2022;13:e863702. https://doi.org/10.3389/fmicb.2022.863702
Jayashree Lakshmi P, Vanmathi Selvi K. Genetic analysis of low-density polyethylene degrading bacteria from plastic dump sites. Indian J Sci Technol. 2021;13(48):4732-4738. https://doi.org/10.17485/IJST/v13i48.2066
Nedi EG, Ebu SM, Somboo M. Identification and molecular characterization of Polyethylene degrading bacteria from garbage dump sites in Adama, Ethiopia. Environ Technol Innovation. 2024;33:e103441. https://doi.org/10.1016/j.eti.2023.103441
Ebu SM, Ray L, Panda AN, Gouda SK. De novo assembly and comparative genome analysis for polyhydroxyalkanoates-producing Bacillus sp. BNPI-92 strain. J Genet Eng Biotechnol. 2023;21(1):e132. https://doi.org/10.1186/s43141-023-00578-7
Sachan RSK, Kumar A, Karnwal A, Paramasivam P, Agrawal A, Ayanie AG. Screening and characterization of PHA producing bacteria from sewage water identifying Bacillus paranthracis RSKS-3 for bioplastic production. BMC Microbiol. 2025;25(1):e136. https://doi.org/10.1186/s12866-025-03841-8
Johnston B, Jiang G, Hill D, et al. The molecular level characterization of biodegradable polymers originated from polyethylene using non-oxygenated polyethylene wax as a carbon source for polyhydroxyalkanoate production. Bioengineering. 2017;4(3):e73. https://doi.org/10.3390/bioengineering4030073
Manoli M-T, Nogales J, Prieto A. Synthetic control of metabolic states in pseudomonas putida by tuning polyhydroxyalkanoate cycle. MBio. 2022;13(1):e01794-21. https://doi.org/10.1128/mbio.01794-21
Arias S, Bassas‐Galia M, Molinari G, Timmis KN. Tight coupling of polymerization and depolymerization of polyhydroxyalkanoates ensures efficient management of carbon resources in Pseudomonas putida. Microb Biotechnol. 2013;6(5):551-563. https://doi.org/10.1111/1751-7915.12040
Wang P, Qiu Y-Q, Chen X-T, Liang X-F, Ren L-H. Metabolomic insights into polyhydroxyalkanoates production by halophilic bacteria with acetic acid as carbon source. Biosci Biotechnol Biochem. 2019;83(10):1955-1963. https://doi.org/10.1080/09168451.2019.1630252
de Eugenio LI, Garci P, Luengo JM, San Roma J, Garci JL. Biochemical evidence that phaZ gene encodes a specific intracellular medium chain length polyhydroxyalkanoate depolymerase in Pseudomonas putida KT2442: characterization of a paradigmatic enzyme. J Biol Chem. 2007;282(7):4951-4962. https://doi.org/10.1074/jbc.M608119200
Di Y, Xia H, Jiao Y, et al. Biodegradation of polyhydroxybutyrate by Pseudomonas sp. DSDY0501 and purification and characterization of polyhydroxybutyrate depolymerase. Biotech. 2019;9:e359. https://doi.org/10.1007/s13205-019-1871-9
Zamanzadeh-Nasrabadi SM, Mohammadiapanah F, Hosseini-Mazinani M, Sarikhan S. Salinity stress endurance of the plants with the aid of bacterial genes. Front Genet. 2023;14:e1049608. https://doi.org/10.3389/fgene.2023.1049608
Copyright (c) 2026 Rafeya Sohail, Rida Batool, Nazia Jamil

This work is licensed under a Creative Commons Attribution 4.0 International License.
Author(s) retain copyright and grant the journal right of first publication with the work simultaneously licensed under a Creative Commons Attribution (CC-BY) 4.0 License that allows others to share the work with an acknowledgment of the work’s authorship and initial publication in this journal.






